|
我們實驗室主要研究細胞分化的過程。在發育過程中,胚胎細胞藉由分化過程來形成各式功能的細胞。分化作用也會在成體中發生,並對成體組織代謝及恆定極為重要。因此,研究細胞分化不只有助於了解及治療因細胞功能不足而產生的先天性疾病,且直接有助於未來在醫學上產生新生細胞來修補受損組織。
我們主要用果蠅這個模式生物來研究在活體中,基因如何調控細胞分化。實驗室中有兩個主要方向:(1)探討神經母細胞的形成過程中,原神經蛋白如何活化下游基因來促進母細胞分化。我們目前著重於探討核內肌動蛋白和原神經蛋白之間的關係。(2)成體幹細胞 (adult stem cell) 自我新生的調控在 維持組織恆定中扮演 重要功能 。自我新生能力太強時, 細胞分化會被抑制。 反之, 過早進入細胞分化會降低幹細胞的自我新生。我們實驗室最近的研究在探討成體中的配子幹細胞如何受到BMP 或其它訊息傳導路徑的影響來維持在自我新生及細胞分化中的協調。
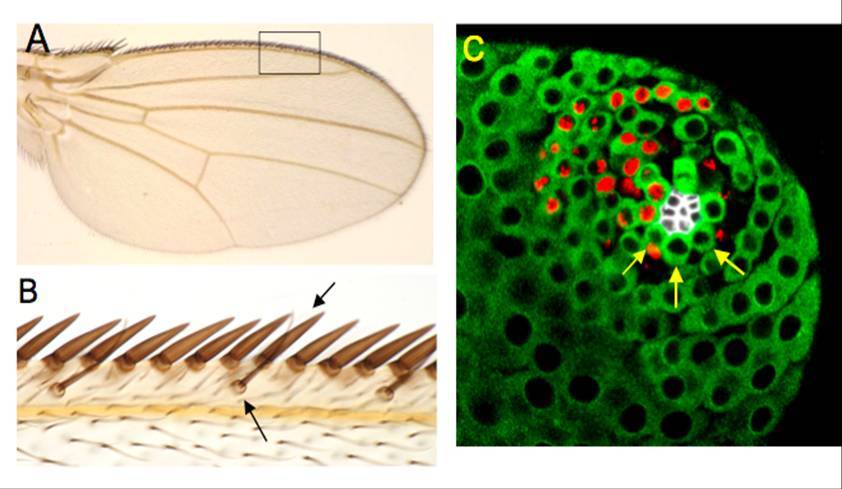
(A)果蠅的翅膀 (B)感覺神經剛毛(箭頭)位於果蠅前翅脈邊緣,負責感覺外界機械性刺激 (C)果蠅成體testis前端,圍繞在hub(中心白色細胞)周圍的細胞為成體配子幹細胞(箭頭)
近五年計劃:
行政院國家科學委員會:
2012/8/1~2015/7/31 探討COP9複雜亞單位對於果蠅蛻皮激素訊息傳遞鏈的抑制機轉
2015/8/1-2018/7/31 Arp6 及 H2Av 在果蠅外感覺器官分佈及發育的功能
2018/8/1-2019/7/31 bHLH 原神經蛋白在神經母細胞中對核醣體的生化合成的調控
長庚醫院:
2013/1/1~2015/12/31 老化果蠅雄性生質幹細胞系統中,smurf扮演抑制細胞過度增生的角色
2016/6/1-2019/5/31 在雄性果蠅性腺中,JNK訊息傳遞鍊調節囊幹細胞恆定的分子機制
|
|
Publications:
- Yun-Ling Hsiao, Hui-Wen Chen, Kuan-Han Chen, Bertrand Chin-Ming Tan, Chia-Hsiang Chen, and Haiwei Pi (2023, February). Actin-related protein 6 facilitates proneural protein-induced gene activation for rapid neural differentiation. Development, accepted. (SCI: 6.862; Rank: 12.8% (5/39) in Developmental Biology). This paper is selected as a “Research Highlight” in Development journal.
- Hsun Li, Hsin-Ho Sung, Yi-Chun Huang, Ying-Ju Cheng, Hsiao-Fong Yeh, Haiwei Pi, Edward Giniger, and Cheng-Ting Chien (2022, Sep). Fringe-positive Golgi outposts unite temporal Furin 2 convertase activity and spatial Delta signal to promote dendritic branch retraction. Cell Reports, 40:111372. (SCI: 9.995; Rank: 17% (33/195) in Cell biology)
- Pei-Yu Wang, Archan Chakraborty, Hsin-Ju Ma, Jhen-Wei Wu, Anna C.-C. Jang, Wei-Cheng Lin1, Hai-Wei Pi, Chau-Ting Yeh, Mei-Ling Cheng, Jau-Song Yu and Li-Mei Pai (2022, Aug). Drosophila CTP synthase regulates collective cell migration by controlling the polarized endocytic cycle. Development, dev200190. (SCI: 6.862; Rank: 12.8% (5/39) in Developmental Biology)
- Yi Chieh Chang, Hsin Tu, To‑Wei Huang, Bo‑Wen Xu & Haiwei Pi* (2020, Jul). Upregulated TNF/Eiger signaling mediates stem cell recovery and tissue homeostasis during nutrient resupply in Drosophila testis. Scientific Reports, 10:11674. (SCI: 4.997; Rank: 25.6% (19/74) in Multidisciplinary Science)
- Kun-Yang Lin, Wen-Der Wang, Chi-Hung Lin, Elham Rastegari, Yu-Han Su, Yu-Tzu Chang, Yung-Feng Liao, Yi-Chieh Chang, Haiwei Pi, Bo-Yi Yu, Shu-Hwa Chen, Chung-Yen Lin, Mei-Yeh Lu, Tsu-Yi Su, Fei-Yang Tzou, Chih-Chiang Chan & Hwei-Jan Hsu (2020, Jun). Piwi reduction in the aged niche eliminates germline stem cells via Toll-GSK3 signaling. Nature Communications, 11:3147. (SCI: 17.694; Rank: 8% in Multidisciplinary Science).
- Yi Chieh Chang, Hsin Tu, Jing-Yi Chen, Ching-Chin Chang, Shu Yuan Yang, Haiwei Pi* (2019, Jul). Reproduction disrupts stem cell homeostasis in testes of aged male Drosophila via an induced microenvironment. PLOS Genetics, 15 (7): e1008062. (SCI: 6.02; Rank: 15.4% in Genetics and Heredity). This paper is featured in PLOS Genetics by an accompanying Perspective article “ Irreversible effects of youthful choices in aged adults”.
- Yang SY*, Chang YC, Wan YH, Whitworth C, Baxter EM, Primus S, Pi H, Van Doren M*. (2017, Aug). Control of a Novel Spermatocyte-Promoting Factor by the Male Germline Sex Determination Factor PHF7 of Drosophila melanogaster. GENETICS, 206(4),1939-1949. (SCI: 4.4; Rank: 32% (56/175) in Genetics and Heredity)
- Chien-Hsiang Wang, Yi-Chun Huang, Pei-Yi Chen, Ying-Ju Cheng, Hsiu-Hua Kao, Haiwei Pi, Cheng-Ting Chien (2017, May). USP5/Leon deubiquitinase confines postsynaptic growth by maintaining ubiquitin homeostasis through Ubiquilin. eLife, 10(6),e26886. (SCI: 8.713; Rank: 8.5% (8/94) in Biology)
- Shih-Han Kao, Chen-Yuan Tseng, Chih-Ling Wan, Yu-Han Su, Chang-Che Hsieh, Haiwei Pi, and Hwei-Jan Hsu (2015, Feb). Aging and insulin signaling differentially control normal and tumorous germline stem cells. Aging Cell, 14(1): 25–34. (SCI: 11.005; Rank: 15.3% (30/195) in Cell Biology)
- Yi-Chun Huang,Yu-Nung Lu,June-Tai Wu,Cheng-Ting Chien, Haiwei Pi* (2014, Nov). The COP9 Signalosome Converts Temporal Hormone Signaling to Spatial Restriction on Neural Competence. PLOS Genetics, 10(11),1-13. (SCI: 6.02; Rank: 15.4% in Genetics and Heredity)
- Yun-Ling Hsiao, Yu-Ju Chen, Yi-Jie Chang, Hsiao-Fong Yeh, Yi-Chun Huang and Haiwei Pi (2014, Jan). Proneural proteins Achaete and Scute associate with nuclear actin to promote formation of external sensory organs. Journal of Cell Science, 127:182-90.
|



